
Free alternatives to commercial electrophysiology software
The recent excitement over quickly developing imaging methods and optogenetics often hides the fact that electrophysiology is still the workhorse of neuroscience. Indeed, most imaging and optogenetics experiments rely on electrophysiology, either throughout the experiment or during characterisation and as control. But even when not combined with any optical methods, electrophysiology is an extremely powerful toolset.
One important aspect of electrophysiology is that it has become possible to set up a rig, or indeed an entire lab, on a very limited budget. Looking through the websites of many suppliers that might not be the first impression a researcher has. Often, amplifiers, digitisers and software are tied into bundles, which are often perceived as expensive. While the simplicity of these integrated systems justifies the price, not all researchers have that level of funds available. But with a bit of effort, it is possible to find lower or zero cost alternatives.
Amplifiers for electrophysiology are available from a wide range of manufacturers, many providing simple but good (e.g. with low noise, fast voltage-clamp) amplifiers. Most of these simpler amplifiers are not computer controlled, which makes them software independent.

Molecular Devices Axon Digidata 1550B
Digitisers have a wide range of prices. There are dedicated electrophysiology digitisers available (e.g. Molecular Device’s Digidata series), in some cases as a single unit with the amplifier (e.g. HEKA’s EPC 10 series). Often these tend to be more expensive than general-purpose digitisers such as those provided by, for example, National Instruments. The advantage of the general-purpose digitisers is that the user can choose a model that fits their experimental needs without paying for functionality they do not need. Another alternative that is often successful is looking through old departmental stock. Many university departments have storage spaces full of ‘outdated’ equipment. Much of this equipment is fully functional, but not supported by the manufacturer’s newest electrophysiology software version anymore.
This is where the third component in an electrophysiological measurement system becomes important: control and measurement software. The commercially available software packages for electrophysiology (e.g. pClamp, Spike2, AxoGraph, PatchMaster) can be expensive, and are in most cases tied to the manufacturer’s own digitiser range; often only supporting recent digitiser models. Again, the price reflects the simplicity of using an integrated solution, but also the technical support these manufacturers tend to offer.
As an alternative to the commercially available software, there is a wide selection of free alternatives available. Most of these programs are written and maintained by neuroscientists, in some cases with strong departmental and funding agency support. Many programs support multiple digitiser families, including ‘outdated’ versions, which can be given a new lease to life by using such software.
The main risks of this type of freeware stem from the fact that they are researcher written: often the functionality is limited to that needed by the authors. Also, most of these programs do not come with technical support. Indeed, quite a few solutions do not even have full documentation. And finally, the development life of the programs tends to be determined by the academic life of the programmer, leading to many solutions that have not been updated in a decade.
Despite the bleak impression that the potential risks might inspire, sifting through the many options available, it is possible to find software packages that have sufficient support from the universities or funding agencies to offer reliability, documentation and in some rare cases even support. In this article, we introduce several free electrophysiology solutions that we believe meet these criteria. We realise this list is not complete, but hope readers will find it a useful starting point for their own exploration.
Before we introduce the different options, we present a quick discussion of features that are common to electrophysiology programs and points to consider when choosing a software package.
Features and functions
Electrophysiology software can have a wide range of features. There is the ‘must have’ functionality, that even the most minimalist software should have, there are capabilities that are ‘nice to have’ and there are additional features that are not necessary but can make life easier.
Fundamental functionality
The most obvious feature that any electrophysiology software needs to have is the ability to acquire data from the amplifier via a digitiser. Along the same lines, the software needs to be able to output voltages via the digitiser, to allow control of the amplifier. Therefore, the software must enable users to configure the digitiser and ideally read information from the amplifier (e.g. filter and gain settings). This configuration can range from rudimentary (e.g. manually assigning digitiser inputs and outputs to the corresponding amplifier inputs and outputs) to nearly full automation, including automatic readout of the amplifier settings over telegraph signals or USB.
A similar level of configuration is needed for digital inputs and outputs (I/O). These are used to trigger both the electrophysiology recording as well external devices such as stimulators that are used in experiments (discussed in detail in “SciMethods: Controlling and Coordinating Experiments in Neurophysiology”). In addition to generic digital I/O, some digitisers have dedicated trigger input ports, to allow for easier configuration. If the software supports these, it makes life a little easier.
The configuration of inputs and outputs is something that is only done when initially setting up a recording rig or when making larger changes, so ease of use here is good, but a more complex setup routine is not necessarily a deal breaker.
More crucial is the last mandatory feature of any electrophysiology software: the ability to generate, store and ideally sequence measurement protocols. Comparable to experimental protocols in molecular biology or biochemistry, a protocol in electrophysiology is a sequence of experimental steps (e.g. depolarize a cell by a defined amount, activate a stimulation electrode, etc.)
Most electrophysiology software provides powerful protocol editors to allow users to set up experiments, ranging from basic assignment of stimulation times or voltage steps, up to more intricate protocols that inject realistic waveforms into cells or deliver varying pulse patterns.
Convenient functionality
Beyond the features that are fundamental to any electrophysiology software, there is a range of features that are convenient to have.
Not just for patch-clamp recordings, a simple means of testing the electrode resistance, seal resistance and basic cell properties (e.g. membrane capacitance, input resistance) is very useful. While it is possible to set up a simple protocol that delivers the required voltage test pulse, a function that runs independently of any protocol and potentially provides a direct read out of the relevant parameters makes life easier.
As mentioned above, generating and managing protocols is function that nearly every user will need from an electrophysiology software. On the other hand, not everyone will need a means of automatically running sequences of protocols. Nonetheless, it is a very convenient function to have for many experiment types. Also useful is the ability to define time intervals between different protocols in a sequence.
Optional functionality
All the features discussed above are ones that should be considered important to have included in the actual electrophysiology software (if they are indeed needed). None of them are easily achieved by using additional software. In contrast to that, there are functions that are commonly handled by additional software, that some electrophysiology software solutions integrate, with the aim of providing tighter integration.
Things such as camera control and image acquisition, sample stage and micromanipulator control, perfusion control or online analysis fall into this category. In general, these are functions that are well addressed by dedicated software, but some experimental workflows benefit from the tight integration with the standard electrophysiology functionality, and thus being able to control everything from a single interface.
Summary on functionality
Ultimately, as usual in experimental research, the best fitting software will be determined by the actual experimental needs. Beyond the fundamental functions, different researchers will have different requirements.
A good example for this is protocol sequencing: Researchers working on plasticity experiments will find sequencing of different protocols (e.g. test stimulation, plasticity induction, test stimulation) convenient, and will commonly require the ability to easily define time intervals between repetitions and different protocols in the sequence. On the other hand, users studying ion channels will want a means of defining sequences of multiphasic voltage steps (e.g. for measuring tail currents) but will be less interested in inter-repetition timing.
Software packages
Below, we introduce several software solutions for electrophysiology. As already mentioned, this list is far from complete and not intended as an endorsement of the individual projects, but as an overview over the different levels of solutions that are available. We start by introducing dedicated electrophysiology software, ranging from software targeting patch-clamp recordings, to more general electrophysiology. After that, we discuss examples of customisable solutions that require more effort from the user but bring the advantage of higher adaptability. Finally, we introduce a few examples of specialized software solutions aimed at specific applications.
Dedicated electrophysiology software
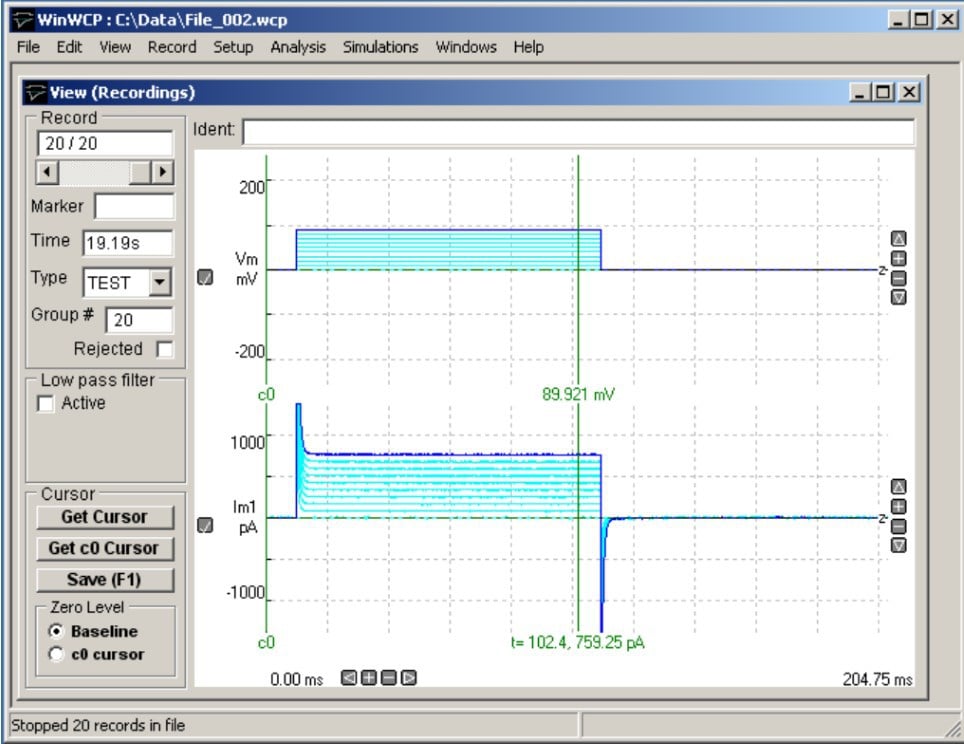
As the name suggests, WinWCP is a Windows software initially developed for whole-cell patch-clamp. Written by Dr John Dempster of Strathclyde University in Glasgow, UK, the software has been continuously under development for over two decades. While the software was developed with patch-clamp recordings in mind, the range of applications it is well-suited for stretches to any microelectrode technique that relies on single electrodes (rather than electrode arrays).
WinWCP is equipped with a powerful protocol editor, that allows the user to generate involved sequences of stimuli within a protocol, but does not allow sequencing of multiple protocols. The one feature that sets WinWCP apart from nearly all other electrophysiology software is the support of nearly all digitiser brands commonly used in patch-clamp electrophysiology, including outdated models of several of these brands.
In addition to the data acquisition capabilities, WinWCP brings a wide range of data analysis routines. This covers a full range of classic cell-physiological methods (e.g. non-stationary noise analysis) as well as basic statistics on responses. Importantly, many of these measurements can be used for online analysis.
In contrast to most other solutions, it is a standalone software, not requiring MATLAB, Igor Pro, or LabVIEW to be installed to run, potentially saving license fees. The possible downside to this is that the software is not easily extendable.
DataPro is an electrophysiology suite continuously developed in Professor Nelson Spruston’s lab for close to twenty years. The development is aimed at patch-clamp electrophysiology but should be applicable to other forms of single electrode recordings (it will support several electrodes, but not electrode arrays). In addition, it offers support for imaging data acquisition using a limited selection of cameras. For data acquisition, it relies on Instrutech ITC 16 and 18 digitisers.
DataPro is implemented in Igor Pro (www.wavemetrics.com), a data analysis and graphing environment often used in patch-clamp labs. Due to the openness of the code, it is possible to add extensions and develop specialised measurement protocols. In addition, the Igor Pro functionality makes implementing online data analysis that runs during the experiment straightforward. This will be of particular interest to researchers performing long-term plasticity experiments.
WaveSurfer is an extensible electrophysiology suite developed by Dr Adam Taylor at Janelia Farm Research Campus. Initially released in 2013, it has strong support from multiple labs at Janelia Farm, all but ensuring future development.
Being open source and written in MATLAB, it is straightforward to add required analysis or visualisation routines, including online analysis. The software is designed to allow such custom routines to be called at defined time point during the acquisition. The result is that WaveSurfer allows closed loop experiments (in 'soft' real time, latency usually <10 ms).
Of particular interest to researchers combining electrophysiology with multiphoton imaging, WaveSurfer can be tightly integrated with Vidrio Technologies’ ScanImage. This integration goes beyond synchronised data acquisition, enabling protocols to be saved, loaded and run in parallel without the need to switch programs.
This tight integration with ScanImage is enabled by WaveSurfer’s reliance on National Instruments X-Series data acquisition devices. The software is not able to acquire data from other manufacturer’s digitisers.
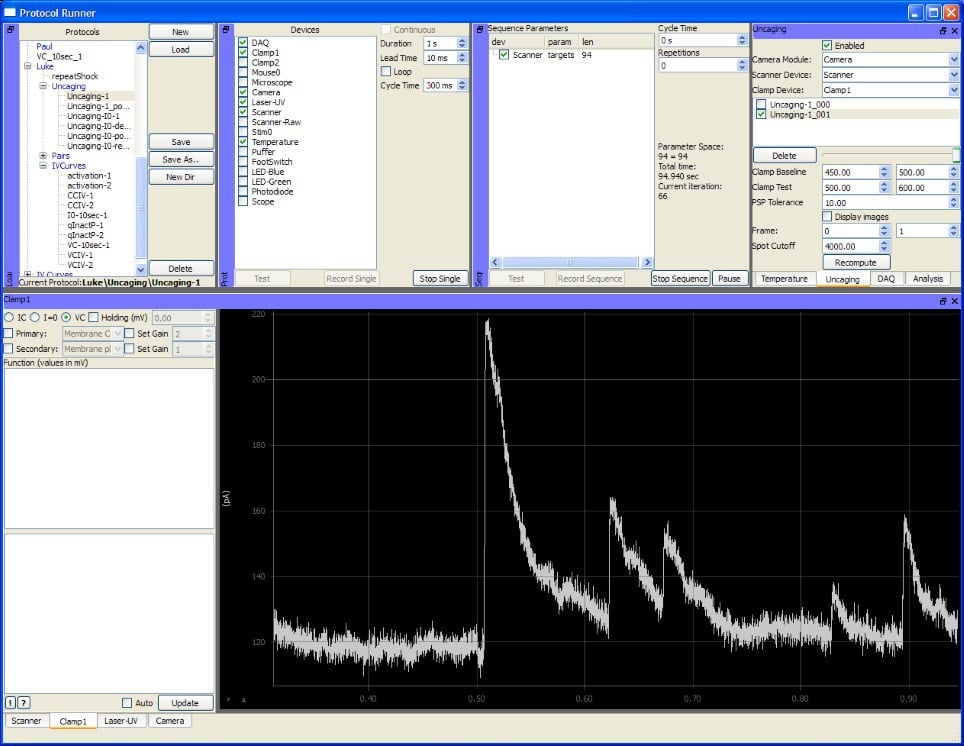
ACQ4 is an extensible general neurophysiology software solution, having been developed by Dr Luke Campagnola at different institutions over the past years. The software is written in Python and open source, allowing for easy extension by the user to suit specific experimental needs. For digitisation, it relies on a wide range of National Instruments hardware.
In addition to electrophysiology, where it focuses mainly on patch-clamp recordings, ACQ4 provides support for controlling micromanipulators, cameras and laser scanners, even allowing for confocal or multiphoton imaging. Combining all these measurement modes, enables users to run involved experiments. The combination of camera and micromanipulator control is designed to allow automation of pipette positioning, to increase the efficiency of patch-clamp experiments with large numbers of electrodes.
Adaptable and customisable solutions
Besides the dedicated electrophysiology solutions introduced above, it is possible to use general data acquisition solutions to acquire electrophysiology data.
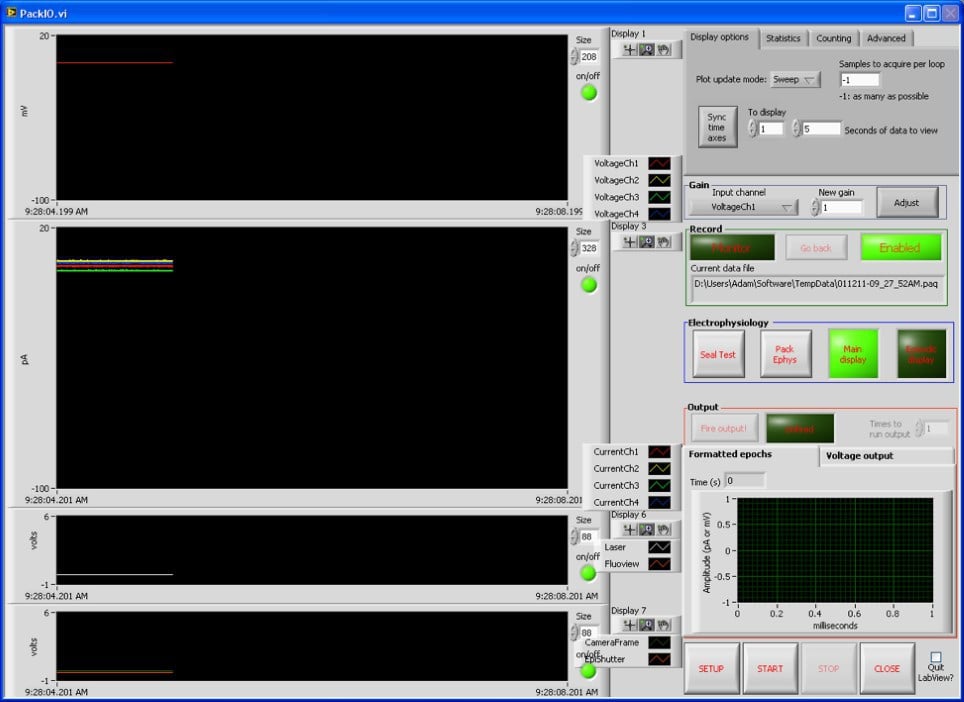
Pack I/O is a general-purpose data acquisition software developed by Dr Adam Packer over the past decade. Developed in LabVIEW, it should be capable of working with most National Instrument data acquisition devices. While the software includes some dedicated modules for patch-clamp electrophysiology, the main aim of the software is to provide researchers with a straightforward means of acquiring and outputting a multitude of analogue and digital signals.
The strength of Pack I/O is the ability to act as a link between different measurement systems, in parallel to acquiring electrophysiological data. For researchers looking to synchronise multiple data streams, this is probably one of the most flexible solutions available.
DIY
One alternative many researchers tend to rule out is writing their own acquisition software. While there is a considerable risk of sinking resources (especially time) into re-inventing the wheel, simple data acquisition programs optimised to solve a clearly defined task have their place. Many digitiser manufacturers provide libraries or programming interfaces to allow easy access to their equipment from the most common programming and data analysis environments (i.e. LabVIEW, MATLAB, Igor Pro and Python).
Special purpose software
A full overview of free electrophysiology software would not be complete without providing some examples of software geared at solving very specific tasks or aimed at specific sets of applications. Below we introduce two examples of specialised software.
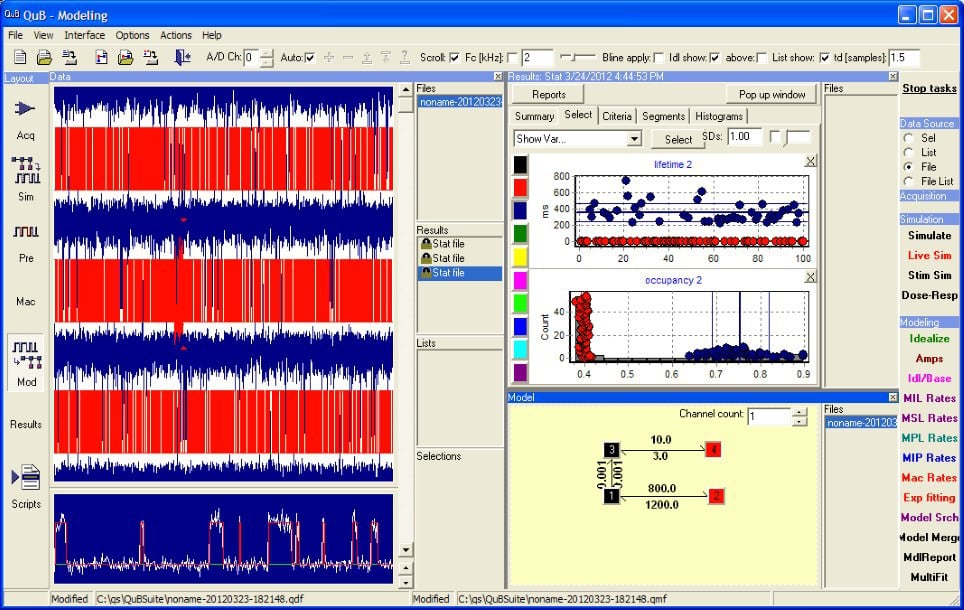
Over the past twenty years, the University of Buffalo developed QuB has evolved from a program for analysing single ion channel recordings by fitting Q-Matrix models, into a full set of tools for single ion channel research. Today, this package includes two data acquisition options (QuB classic and QUBIO), optimized for recordings from single ion channels. In addition, the software includes a simulation engine.
Today, there are QuB development branches that include modules that generate a 3D model of the recording environment and use machine vision routines interacting with micromanipulator controls to automatically position recording electrodes. In addition, this branch (‘The MLab Edition’) has been extended with features for dynamic clamp experiments, network simulations, and more, making it to a true workhorse for cell physiologists.
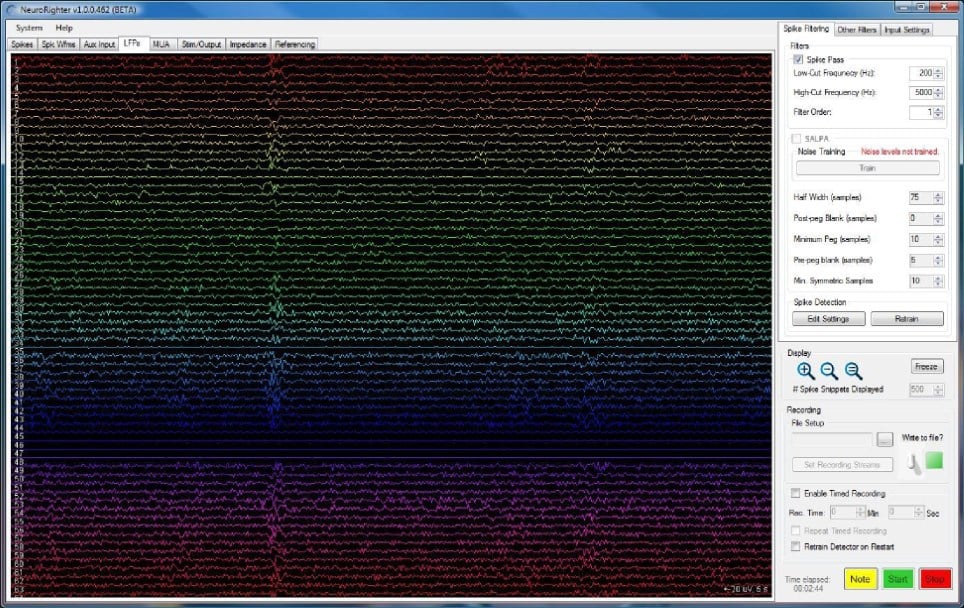
Most software discussed so far is mainly aimed at researchers using single microelectrodes, mostly for patch-clamp recordings. Some of the more general solutions presented above could be configured to work with microelectrode arrays, but in most cases, this was never the idea. In contrast to this, NeuroRighter is a software dedicated to working with microelectrode arrays. Developed by the Potter lab at Georgia Tech, NeuroRighter is designed to allow closed-loop experiments using Multichannel Systems recording hardware and National Instruments digiters. It allows for recording and stimulation and offers some initial data analysis functions (e.g. spike sorting).
Conclusion
In this article, we provided a first introduction to several free electrophysiology software solutions. Confronted with the sheer mass of such programs, we handpicked some examples that we believe are powerful, reliable and future-proof. To allow the novice user to evaluate other options not discussed here, we provide an overview over functions we believe to be of importance. Obviously, the planned experiments will determine which functions are needed, and which are not. To help establish a first idea of this, we use these criteria to develop an idea of what types of work the different packages are suited for.
We hope to have given the reader a first understanding of what can be achieved with free electrophysiological software. While we believe that these solutions have advantages in situations where funding or equipment are limited, it does need to be said that nearly all these solutions do not have any software support and in some cases only basic manuals. Thus, for a researcher looking for a quick and easy start into electrophysiology, the available commercial packages (e.g. from Molecular Devices, HEKA, or Cambridge Electronic Design) are a reasonable option.
Scientifica SliceScope Pro 1000
The SliceScope Pro 1000 is Scientifica's most cost effective complete electrophysiology rig. This package consists of the SliceScope microscope, motorised XY translation stage, two PatchStar manipulators and adjustable Post & Platform mounting systems for the manipulators and sample.




)